Forschungsprojekt MEDVIS 3D
Blutflusssimulation im Gehirn auf Basis medizinischer Bilddaten.
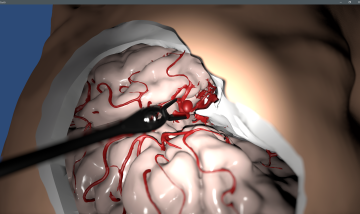
Das Ziel des Forschungsprojekts MEDVIS 3D, das gemeinsam mit dem Kepler Universitätsklinikum Linz (Med Campus III, vormals AKH Linz, und Neuromed Campus, vormals LNK Wagner-Jauregg) entwickelt wurde, ist ein universelles Softwarewerkzeug zur einfachen und schnellen Rekonstruktion von Aneurysmen aus medizinischen Bilddaten (MR, CT, etc.). Basierend auf den Schichtdaten dieser bildgebenden Verfahren kann die Software das aufgenommene Volumen dreidimensional rekonstruieren und direkt visualisieren. Das System erkennt Areale, in denen eine krankhafte Ausdehnung der Gefäßwand (Aneurysma) vorliegt und kann verschiedene Maßzahlen wie Durchmesser, Volumen und Schnittflächen berechnen. Dies kann in Echtzeit durch direkte Interaktion von dem*der Mediziner*in weiter verfeinert werden. Dadurch wird die Diagnose unterstützt sowie eine qualitative Bewertung des Therapiefortschritts ermöglicht.
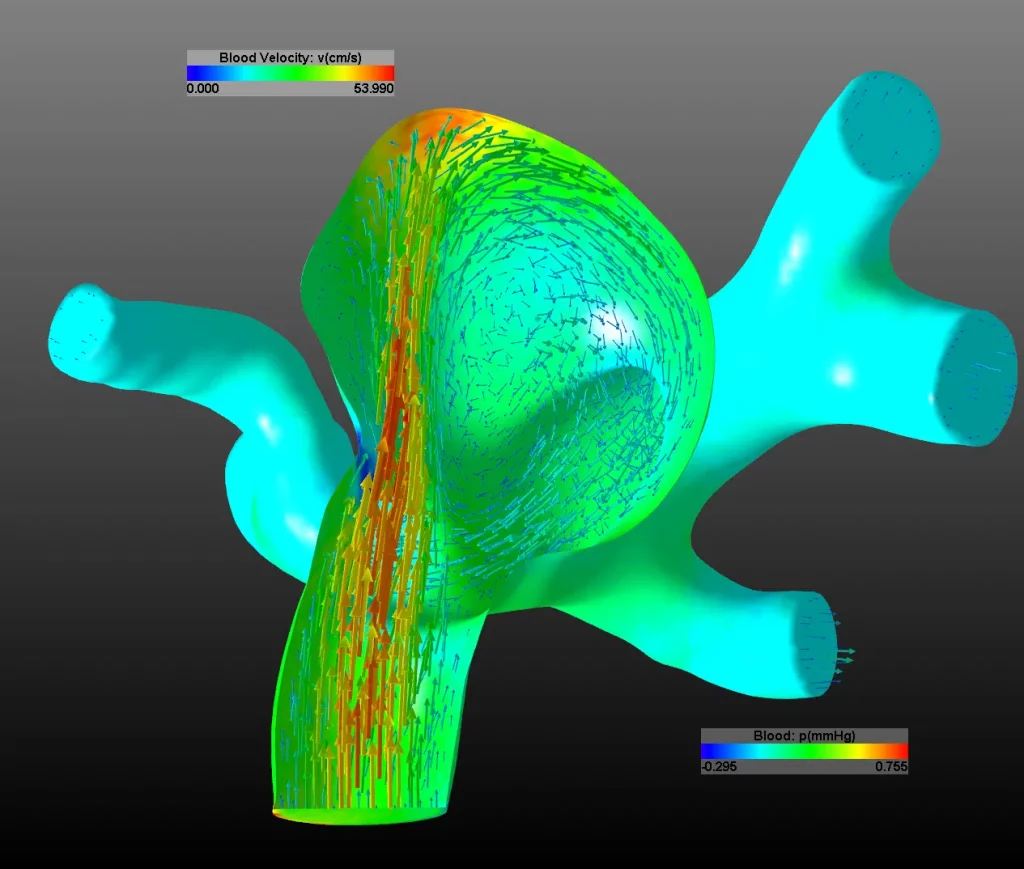
Die rekonstruierten 3D-Geometriedaten können auf Knopfdruck in ein Computersimulationsmodell integriert werden. Mittels der Finiten Elemente Methode (FEM) kann der Blutfluss im Gehirn simuliert werden. Die aufwendigen numerischen Lösungsverfahren werden mittels Algebraic MultiGrid Verfahren (AMG), High Performance Computing (HPC) und GPU Programmierung hardwarebeschleunigt berechnet. Je nach Komplexität der Gefäßstruktur dauert ein Simulationszyklus wenige Minuten bis einige Stunden. Die Simulation kann Geschwindigkeits- und Druckfelder sowie Oberflächenspannungen an den Gefäßwänden berechnen und auch die Ausdehnung der Gefäße durch den Herzschlag berücksichtigen. Damit können das Risiko für eine Hirnblutung sowie adäquate Therapiemaßnahmen ermittelt werden. Gemeinsam mit Ärzt*innen wurde das System im klinischen Einsatz evaluiert und die ermittelten Ergebnisse in einer zentralen Datenbank abgelegt. Dadurch können die mechanischen Prozesse bei der Entstehung und dem Wachstum von Aneurysmen weiter erforscht werden. Eine klinisch einsetzbare Version von MEDVIS 3D ist seit 2013 verfügbar
Diese Forschungsarbeit wurde mit Mitteln des Landes Oberösterreich, der GESPAG (nun OÖG) sowie mit Fördermitteln der FFG finanziert.



Ansprechperson

Dr. Michael Giretzlehner
Head of Unit Medical Informatics